Hamiltonian Paths Demo in Fortran #
1 Simulation and Analysis of Graph Path Selection #
Graph theory serves as a fundamental framework for modeling and understanding various systems across scientific domains. This study presents a detailed computational simulation implemented in Fortran, focusing on the manipulation, selection, and analysis of paths and individual entities within a graph. The simulation involves stepwise operations, including path creation, criteria-based selection, and the identification of Hamiltonian paths. Through this process, the efficacy of Fortran in managing graph structures is explored and analyzed.
2 Fortran #
One of the primary reasons for opting for Fortran was its robustness and reliability in handling numerical simulations. Additionally, Fortran’s performance in handling array-based operations and its ability to optimize code for high performance were crucial factors. Its speed and efficiency in managing computational tasks, especially those involving heavy mathematical computations, were significant advantages for this simulation.
3 Goal and Strategy #
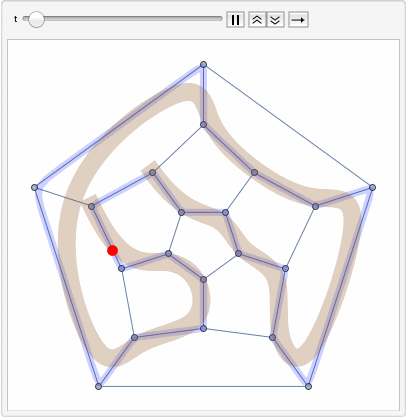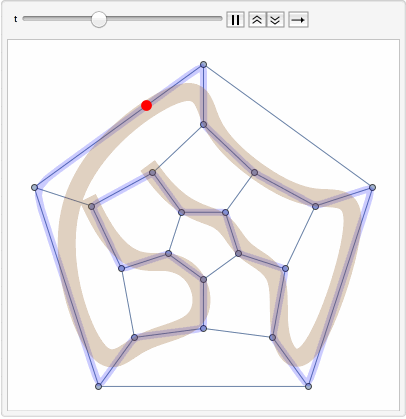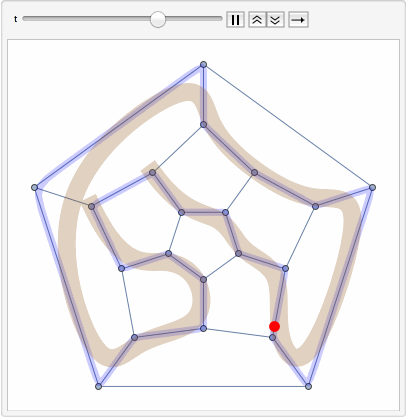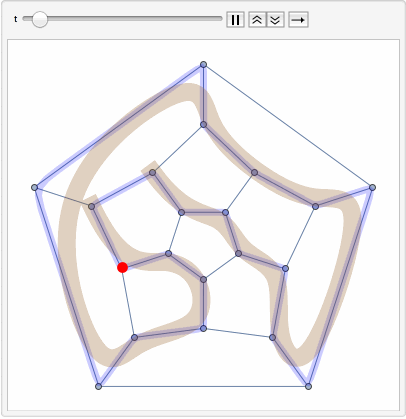
The primary goal is to identify Hamiltonian paths, which traverse all nodes in a graph exactly once, using a combination of path concatenation, selection, and filtering operations within the soup.

The steps of the program involve preparing random edges between nodes, populating the soup with individuals, and visualizing these edges. It also involves path concatenation, selection based on specific criteria, and identification of Hamiltonian paths.
Core functionalities include linked list operations for path representation, concatenation, and selection. Filtering criteria are applied to identify potential Hamiltonian paths within the graph structure.
3.1. Methodology & Code Implementation #
This project introduces a simulator program that encompasses various graph operations within a modular framework. The program is designed to facilitate the analysis and manipulation of graph structures represented as a collection of paths within a soup. The simulator leverages three essential modules: msoup, mind, and mpath, each contributing distinctive functionalities towards simulating, filtering, and identifying specific path attributes within the graph structure.
tip
My Github Repo (Disclaimer: There is a C++ function!)
3.2. Modules #
3.2.1. msoup #
The msoup module in this Fortran implementation serves as a robust framework for managing a collection of individuals, each representing a specific path within a graph structure. At its core, this module encapsulates a soup type, acting as a container for these individuals, and a node type, serving as the building block for storing individual references.
Within the msoup module, the soup type comprises pointers to the first and last individuals in the collection (fst and lst), alongside an integer count (ln) indicating the number of individuals present. This structure offers an organized and efficient means to handle and manipulate these individuals.
type, public :: soup
private
type(node), pointer :: fst ! Points to the first individual of path
type(node), pointer :: lst ! Points to the last individual of path
integer :: ln ! Number of vertices (length) of path
end type soup
Individuals themselves are represented as instances of the node type. Each node stores a reference to an individual (ind) and a pointer to the next individual (next) within the collection. This linked structure allows for easy addition, removal, and traversal of individuals within the soup.
Functionally, the msoup module provides essential procedures:
newS()initializes an empty soup, setting pointers to null() and the individual count to zero.
function newS() result(s)
type(soup) :: s
s%fst => null() ! Set the first individual pointer to null
s%lst => null() ! Set the last individual pointer to null
s%ln = 0 ! Initialize the count of individuals to zero
end function newS
insertS()adds an individual to the soup by creating a new node, associating it with the provided individual, and updating pointers accordingly.
subroutine insertS(i, s)
type(ind), intent(in) :: i
type(soup), intent(inout) :: s
type(node), pointer :: aux
allocate(aux)
aux%ind = i
if (s%ln > 0) then
s%lst%next => aux ! Link the new node to the last node
else
s%fst => aux ! Update the first node if the soup is empty
end if
s%lst => aux ! Update the last node to the newly added node
s%ln = s%ln + 1 ! Increment the count of individuals
end subroutine insertS
important
To acess the 1st, last and count of individuals in the soup:
! Accessing the first individual in the soup individual_1 = sp%fst%ind ! Accessing the last individual in the soup individual_last = sp%lst%ind ! Accessing the count of individuals in the soup number_of_individuals = sp%ln
nextS()identifies pairs of individuals with compatible paths, computing the shortest distance between them, and returning these individuals along with a success indicator.takeS()removes a specified individual from the soup, adjusting pointers and deallocating the node.an_indS()retrieves a random individual from the soup, likely accessing the first individual’s path.emptyS()checks if the soup is empty by examining the association of pointers.sizeS()returns the count of individuals in the soup.showS()displays details of all individuals in the soup, including their coordinates and paths.
3.2.2. mind #
The mind module handles the characteristics of individual entities (ind). It encapsulates their spatial coordinates and associated paths. This module includes functionalities for creating individuals, accessing their coordinates, calculating distances between them, and determining their equality. This ind type comprises attributes for a path (w) and three real-numbered coordinates (x, y, z).
use mind
type(path) :: myPath
type(ind) :: myPoint
! Assume initialization of myPath and coordinates for myPoint
call makeI(myPath, 1.0, 2.0, 3.0, myPoint)
! Retrieving coordinates
real :: x_coord, y_coord, z_coord
x_coord = xposI(myPoint)
y_coord = yposI(myPoint)
z_coord = zposI(myPoint)
As visible, within this module, there are several functions and subroutines:
makeI: This subroutine creates an ind object with specified coordinates and a path.
subroutine makeI(w, x, y, z, i)
type(path), intent(in) :: w
real, intent(in) :: x, y, z
type(ind), intent(out) :: i
i%w = w
i%x = x
i%y = y
i%z = z
end subroutine makeI
pathI: Extracts the path attribute from an ind object.xposI,yposI,zposI: These functions retrieve the x, y, and z coordinates from an ind object.distI: Computes the Euclidean distance between two ind objects using their coordinates.
important
This is only the Euclidean distance!
function distI(i1, i2) result(r) type(ind), intent(in) :: i1, i2 real :: r r = sqrt(((i1%x - i2%x)**2) + ((i1%y - i2%y)**2) + ((i1%z - i2%z)**2)) end function distI
ind_eqI: This function checks if two ind objects have the same coordinates.
important
function ind_eqI(i1, i2) result(b) type(ind), intent(in) :: i1, i2 logical :: b if ((xposI(i1) == xposI(i2)) .and. (yposI(i1) == yposI(i2)) .and. (zposI(i1) == zposI(i2))) then b = .true. else b = .false. end if end function ind_eqINote: As previously mentioned, this function compares if objects (
i1andi2) have identical coordinates. It compares coordinates usingxposI,yposIandzposIfunctions and returns a logical value.
note
These implementations illustrate how the functions extract coordinates (xposI,yposI,zposI), calculate distances between points (distI), and determine if points share identical coordinates (ind_eqI).
3.2.3 mpath #
The mpath module in Fortran is dedicated to managing paths using linked lists, represented by the path and node types. The path type contains pointers to the first and last elements of the path (fst, lst) and keeps track of the path’s length (ln). On the other hand, each node represents an element in the path, storing an integer value and a pointer to the next node. As visible, within this module, there are several functions and subroutines:
newPcreates an empty path by setting pointers to null and initializing the length as zero.enterPappends an element to the end of a path, updating pointers accordingly.
subroutine enterP(x, w)
integer, intent(in) :: x
type(path), intent(inout) :: w
type(node), pointer :: aux
allocate(aux)
aux%value = x
if (w%ln > 0) then
w%lst%next => aux
else
w%fst => aux
end if
w%lst => aux
w%ln = w%ln + 1
end subroutine enterP
note
In turn, it allocates memory for a new node, assigns the value x to it, updates pointers, and increments the path length.
revPcopies a specified number of elements from one path to another in reverse order.shortenPremoves the first element from a path, deallocating memory and updating pointers.makePcreates a path with two specified elements by utilizing newP and enterP.firstPandlastPretrieve the first and last elements of a path, respectively, handling empty path conditions.lengthPreturns the length of a path by accessing the ln attribute.compPchecks if the last element of one path matches the first element of another path.gluePmerges two compatible paths by linking their nodes, creating a new path.crossesPdetermines if a specified value exists within a path by traversing nodes and comparing values.copyPduplicates the contents of a path to another in reverse order using revP.deletePrecursively removes elements of a path until it becomes empty, deallocating nodes.showPdisplays the elements of a path by reversing it and printing its values.
subroutine showP(w)
type(path), intent(in) :: w
type(path) :: r
integer :: i
integer, dimension(w%ln) :: aux
r = newP()
call revP(w, r, w%ln)
do i = 1, w%ln
aux(i) = firstP(r)
if (r%ln > 1) then
call shortenP(r)
end if
end do
print *, aux
end subroutine showP
warning
These functionalities handle path creation, manipulation, comparison, concatenation, element retrieval, deletion, and display efficiently using the linked list structure.
3.3. Simulator #
The simulation process within this program is orchestrated across several distinct steps, each playing a crucial role in analyzing, manipulating, and presenting pathways within a given graph structure.
- #1. Initialization & Graph Preparation: This initial step involves setting up the simulation environment and creating the initial
soup, which holds individual nodes representing edges within the graph.
tip
Key Actions:
- Parameter Setup: Initializes user-defined parameters such as the number of nodes (
no), edges (s), and other simulation variables (ka,nu).- Node Creation: Generates individual nodes representing edges, assigning them random coordinates and attributes within the soup.
- #2. Hybridization & Path Concatenation: Concatenates compatible paths, creating new paths by merging individual paths from the soup, fostering connections in the graph.
do c6=1, nu
call nextS(sp, i1, i2, sb)
call pathI(i1, w1)
call pathI(i2, w2)
call glueP(w1, w2, w3)
...
end do
tip
Key Actions:
- Path Merging: Combines compatible paths from individual nodes to create longer, concatenated paths.
- Creation of New Paths: Generates new paths representing connected segments within the graph by merging compatible paths.
- #3. Path Selection & Filtering: Selects specific paths based on defined criteria, filtering out paths that do not meet user-defined conditions.
do c9=1, no
eln = eln + 1
do c10=1, sizeS(sp3)
call an_indS(sp3, i6)
call pathI(i6, w6)
call crossesP(w6, eln, ib1)
...
end do
end do
tip
Key Actions:
- Selection Criteria: Filters paths based on criteria such as starting and ending nodes, path length, or nodes traversed.
- Soup Reorganization: Removes incompatible paths from the soup while organizing selected paths for further analysis.
- #4. Hamiltonian Path Identification: Identifies and presents Hamiltonian paths, which are paths that traverse all nodes in the graph.
tip
Key Actions:
- Traversal Analysis: Analyzes filtered paths to identify those that traverse all nodes in the graph.
- Presentation of Results: Displays identified Hamiltonian paths for comprehensive graph exploration and analysis.
In the end, the simulation aids in comprehending how different entities or nodes within a network are interconnected. By identifying paths and relationships between nodes, it offers insights into how information, resources, or influence flow through a network. It facilitates the discovery of optimal paths that traverse all nodes or specific ones. This capability is invaluable for planning efficient routes, optimizing processes, or understanding the most effective pathways within a system.
This project served me as an educational tool, offering a foundational understanding of network theory, paths, and connections – providing a hands-on approach to learning about complex systems and their underlying structures.
4 Discussion and Efficiency #
This study represented the efficacy of Fortran in computational simulations for graph path selection and analysis. The utilization of modules (msoup, mind, and mpath) showcases how Fortran manages individual entities, paths, and complex graph structures efficiently. The simulation steps, from initialization to the identification of Hamiltonian paths, illustrate the algorithmic flexibility of Fortran in traversing and analyzing graphs.
The significance of criteria-based path selection is evident in this study. It serves as a fundamental aspect in uncovering specific graph properties, such as Hamiltonian paths. The discussions around computational efficiency and the impact of different criteria on path selection highlight the practical relevance of these simulations in diverse scientific domains.
example
Moving forward, several promising directions emerge for advancing this research. Algorithmic improvements hold potential for enhancing computational efficiency, especially when dealing with larger and more intricate graphs. Optimizing path selection methodologies could significantly expedite the analysis process.
Investigating parallel computing techniques stands as another avenue to explore, leveraging Fortran’s capabilities for faster processing of extensive graph structures and enabling more extensive analyses.
Lastly, integrating machine learning techniques into the analysis pipeline could refine criteria determination, potentially improving accuracy and adaptability in path selection methodologies.